Why you should incorporate spatial biology into your research
Spatial biology is a rapidly growing field. It is important because cellular location and context matter when it comes to understanding disease progression, identifying biomarkers, developing therapeutics and monitoring therapeutic response. Recent research consistently shows that spatial mapping of the tissue architecture and cellular landscape can resolve correlated cell states and signaling networks that underlie organ function and pathology.
What is spatial biology?
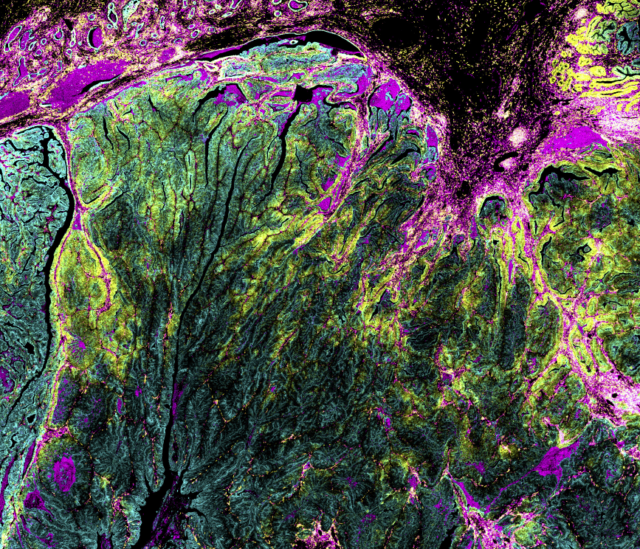
Spatial biology explores how cells and molecules are distributed within tissues and how they interact in context. It involves investigation of the three-dimensional organization of biological systems, which highlights mechanisms and functions not captured, or fully understood, without their structural context. This is particularly important because tissues are highly heterogeneous, composed of distinct cell types organized in ways that support their survival.
In oncology, solid tumors are dynamic ecosystems, containing malignant and non-malignant cells – including cancer, stromal and immune cells – that change over time and space in response to intrinsic and extrinsic factors. Exploring cell phenotypes, spatial relationships and cell-cell interactions in the tissue microenvironment provides insight into biological processes and disease pathogenesis.
Generating spatial information at the single-cell level is key to assessing or predicting cellular activity and determining how specific pathways operate in certain environments. Researchers can also use spatial biology techniques to study cell differentiation, immunosuppression in tumors, responses to new therapies, supporting advances that improve patient outcomes.
Nature Methods names spatial proteomics Method of the Year 2024
Spatial proteomics is integral to spatial biology applications, and is the basis for Imaging Mass Cytometry technology. Bringing spatial proteomics to the forefront highlights the importance of understanding complex biology in a spatial context, and how proteomics in particular offers a unique look into cell function and interactions that create a healthy tissue state or into the cause of dysregulation that results in disease.
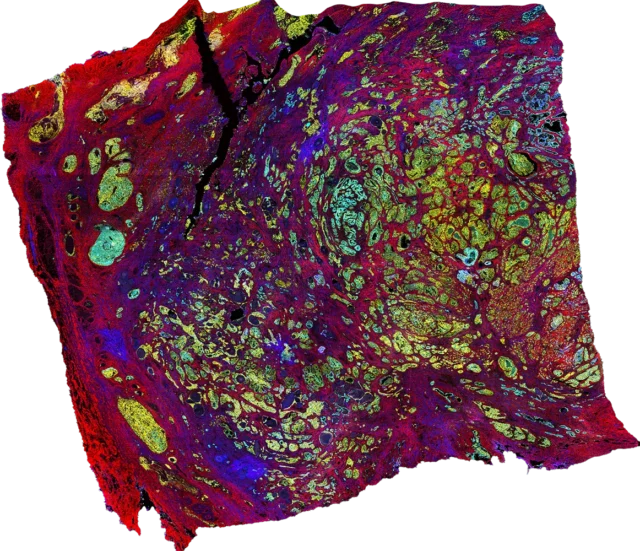
Spatial omics technologies at the single-cell level
Spatial omics technologies vary in spatial resolution, throughput capability and multiplexing capacity and have powered our knowledge of biology, with multiple methods being used to characterize tissues. See some of these techniques in more detail:
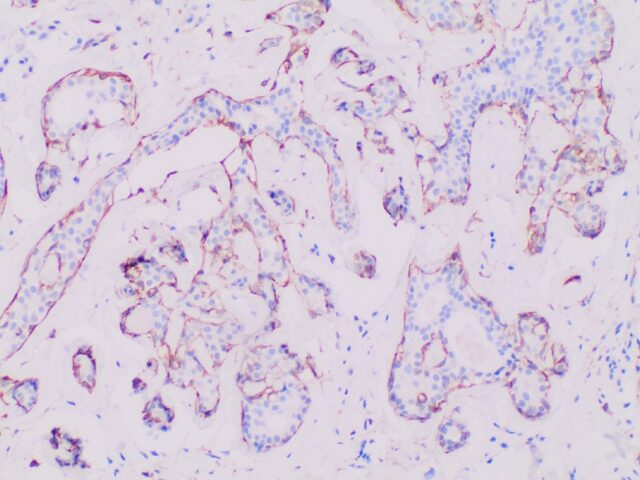
Immunohistochemistry (IHC)
Traditional pathology methods such as immunohistochemistry are the gold standard in characterizing tissue morphology and confirming target molecule expression. When combined with genetic, RNA transcript and protein information, they can be used to decipher the molecular pathways that correlate to an observed phenotype.
Bulk and RNA-seq analysis
Conventional bulk molecular biology techniques including RNA sequencing provide information on the overall RNA content or behavior of cell populations, but do not convey spatial information. These techniques can be more meaningful if spatial context is included with the data.
Multiplex imaging
Instead of analyzing single cells in suspension, multiplex imaging technologies such as IMC™ result in high-dimensional analysis that can substantially broaden what can be discerned from one tissue sample. IMC reconstructs images from tissue sections scanned at single-cell resolution, with the specificity and high content of mass spectrometry.
Genomics, transcriptomics, proteomics and other omics approaches typically focus on generating data that will lead to specific knowledge about a particular question. Integration of the omics fields promotes a shift in strategy to support better understanding of biological systems as a whole and paves the way for advancement into areas such as precision medicine.
For example, cell state and behavior can be influenced by factors such as genetic aberrations contributing to tumor progression. Quantifying these factors to accurately measure the spatial location of genomic sequences together with phenotypic readouts enables analysis of distinct cell types and their copy number alterations. This type of multi-modal approach that combines spatial genomics with cell phenotyping provides a comprehensive way to identify factors contributing to gene expression changes in the tumor microenvironment.

Hartland Jackson, PhD, an investigator at the Lunenfeld-Tanenbaum Research Institute, Sinai Health, aims to better understand cancer at the single-cell level by investigating how the tumor microenvironment drives disease progression and therapeutic resistance. Before starting his own lab, Jackson worked with the team that pioneered high-plex single-cell analysis with IMC™ technology while completing his postdoc in the Bodenmiller Lab.
One technology with multi-omic capabilities
Detection of RNA and proteins in the same cells enables a more complete picture than from transcriptomic data alone. Since some spatial transcriptomic platforms preserve tissue integrity and protein epitopes, it is possible to perform IMC analysis on the same tissue section. A novel workflow combining spatial transcriptomics and the Hyperion XTi Imaging System enables the localization of both transcripts and proteins on a single slide, providing valuable insights into their regulation and function in tissue microenvironments.
Uniting the power of spatial transcriptomics and proteomics:
- Bridges the gap between gene expression and functional protein output, revealing the full molecular phenotype of cells in situ
- Improves cell type and state resolution, especially when transcriptomic and proteomic markers differ in specificity or abundance
- Uncovers post-transcriptional regulation, such as differing mRNA and protein levels, due to translational control or protein stability
- Enhances spatial context, allowing more accurate mapping of cell-to-cell interactions, signaling pathways and tissue architecture
- Strengthens biomarker discovery and therapeutic targeting, integrating upstream (RNA) and downstream (protein) molecular data
Supporting the move toward personalized medicine
Spatial biology can play an integral role in the progression from discovery through translational research and into the clinic. By identifying underlying mechanisms of cell behavior and responses to therapeutic interventions, spatial biology tools can be used to understand the implications of cell heterogeneity on patient outcomes.
From broad cellular landscape characterization to granular analysis of cell-cell interactions, IMC technology has been used to transform quantitative spatial data into knowledge to answer complex research questions. For example, IMC platforms can help uncover biological signatures that may lead to the next generation of biomarkers important to diagnostic and therapeutic uses.
Whole slide imaging and selection of regions of interest
Depending on the application of spatial biology, researchers might choose between imaging an entire slide or selecting smaller regions of interest. What’s the difference?
Whole slide imaging allows the visualization of the overall heterogeneity of a tissue sample, reveals structural variations across the sample and can assist in pinpointing a focus area of activity.
Regions of interest are a common approach to multiplexed imaging, enabling a deep dive into a smaller section of the tissue for single cell and even subcellular analysis.
View examples of workflows using one or combining both of these approaches:
Applications of spatial biology
Researchers and clinicians apply insights derived from IMC technology to the study of disease, drug development programs and precision medicine.
Use of Imaging Mass Cytometry for spatial analysis
IMC technology facilitates the translation of spatial data to the clinic. Without challenges of autofluorescence, it is the trusted technology to generate images with precision and clarity.
How do IMC systems generate high-quality images?
High-dimensional spatial proteomic technologies utilize labeled antibodies to map the tissue microenvironment. IMC technology combines cytometry by time-of-flight (on which CyTOF™ systems are based) with metal-tagged antibodies to generate spatial biology information. The technique uses a standard immunohistochemistry workflow.
Data analysis of the tissue or tumor microenvironment allows for the phenotyping and quantification of key cell types, the visualization of alterations in tissue architecture and the investigation of complex events at the cellular level. With a robust and reliable workflow, IMC platforms allow for the quick generation of new insights into the pathophysiology of disease.
40 slides. 40-plus markers. 24 hours.
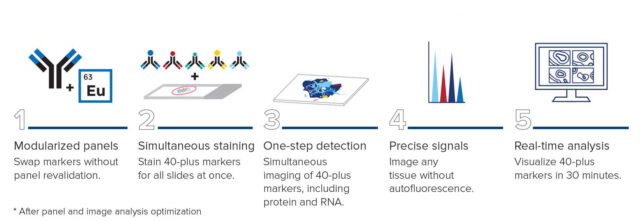
Fast-forward spatial biology. New innovations in the IMC workflow make it possible for researchers to visualize 40-plus markers of the entire tissue in 20 minutes – taking spatial proteomics to new limits.
Imaging Mass Cytometry technology enables ultrafast multiplexed imaging using a unique one-step detection workflow. IMC systems image 40-plus markers simultaneously without time-consuming cyclic protocols or challenges of autofluorescence.

Jana Fischer, PHD
Co-Founder and CEO Navignostics
Hear how the team at Navignostics is translating spatial insights into the clinic by leveraging spatial proteomics into reporting within 72 hours of sample receipt, revolutionizing treatment decisions for each cancer patient.
Frequently asked questions
Spatial biology is the study of how individual cells fit within the context of a tissue, what their environment says about their behavior, where they are and why they are there. The term includes spatial transcriptomics, spatial genomics, spatial proteomics, spatial profiling and spatial omics. The significance of spatial biology in terms of life sciences and biomedical research is its power to expose how cellular interactions determine the tissue structure and spatial organization within the human body and those implications in disease pathogenesis.
IMC technology combines cytometry by time-of-flight with high-resolution laser ablation and metal-tagged antibodies to generate spatial biology information. Tissue sections are stained with panels of antibodies tagged with metal isotopes and scanned, generating plumes of the metal isotopes that are carried to a mass cytometer for simultaneous analysis. Sequential scanning of 1 µm spots yields a high-dimensional map of target proteins in a tissue or region of interest.
The Hyperion™ XTi Imaging System is powered by Imaging Mass Cytometry technology with a simple stain-image-analyze workflow, creating the ultimate platform for multiplex imaging. The Hyperion XTi Imaging System provides the fastest and most reliable tissue imaging workflow for clear and precise spatial analysis.
The use of metal instead of fluorescent tags decreases crosstalk between channels, enabling high-content analysis in a single scan at subcellular resolution. Since IMC technology leverages time-of-flight mass spectrometry, it has the unique capability to simultaneously stain, acquire and analyze the markers of interest on a tissue section without interference from autofluorescent tissues or management of spectral overlap either in panel design or image analysis.
Histology and conventional pathology methods have been central in characterizing tissue morphology, which is then combined with genomic, transcriptomic and proteomic data to determine the molecular programming that correlates to an observed phenotype. Spatial biology techniques provide an in situ molecular characterization for a single-cell map of the complex tissue microenvironment.
Bulk analysis (such as RNA-seq) and single-cell analysis (such as flow cytometry or single-cell sequencing) provide information on either the overall RNA or protein content or behavior of single cells in a suspension. Combining this data with the spatial information of individual cells allows researchers to accurately assess the complex phenotypes and spatial interactions in the tissue microenvironment.
Single-cell analysis by IMC platforms reveals cell phenotypes in context of a particular tissue providing insights into pathological processes. For example, using IMC systems to characterize cellular features, such as immune and stromal cells, of a tumor microenvironment could demonstrate clinical relevance by linking to prognosis.
Notably, IMC technology can be applied to paraffin-embedded tissue sections, allowing its application to the retrospective analysis of patient cohorts. Groups such as the Tumor Profiler (TuPro) Consortium, leading the Tumor Profiler Study, are helping push boundaries by taking an integrative approach to in-depth tumor profiling to support clinical decision making. With efforts to advance multi-omic data integration and improve data analysis solutions, the potential opportunity for IMC technology to impact personalized medicine is becoming increasingly clear.
As an example of taking spatial biology to the clinic, learn how Jana Fischer, PhD, and her company, Navignostics AG, aim to standardize the use of spatial biology data in better informing the most effective treatment decisions for patients, work that could be pivotal in the fight against cancer. Read more here.
Spatial profiling techniques enable the study of cell activity in the context of a tissue microenvironment. By classifying both the transcriptomic and proteomic signatures of a microenvironment, such as a tumor and the associated immune and stromal cells, spatial profiling facilitates evaluation of cell heterogeneity, details the evolution of a tumor and analyzes interactions between each tumor cell and its microenvironment. This provides the information required for correlating clinically relevant features in immuno-oncology, for example.
Innovations in imaging technologies allowing automation (emphasizing consistency, reproducibility and throughput), scalability (improvements on scan times and number of slides processed per day) and even 3D analysis generated from image sets, alongside the development of standardized biomarkers, are further pushing the boundaries of spatial biology into the clinic.
